




ErbB-4 (phospho Tyr1284) rabbit pAb
 One-click to copy product information
One-click to copy product information$148.00/50µL $248.00/100µL
| 50 µL | $148.00 |
| 100 µL | $248.00 |
Overview
| Product name: | ErbB-4 (phospho Tyr1284) rabbit pAb |
| Reactivity: | Human;Mouse;Rat |
| Alternative Names: | ERBB4; HER4; Receptor tyrosine-protein kinase erbB-4; Proto-oncogene-like protein c-ErbB-4; Tyrosine kinase-type cell surface receptor HER4; p180erbB4 |
| Source: | Rabbit |
| Dilutions: | Western Blot: 1/500 - 1/2000. Immunohistochemistry: 1/100 - 1/300. Immunofluorescence: 1/200 - 1/1000. ELISA: 1/10000. Not yet tested in other applications. |
| Immunogen: | The antiserum was produced against synthesized peptide derived from human HER4 around the phosphorylation site of Tyr1284. AA range:1250-1299 |
| Storage: | -20°C/1 year |
| Clonality: | Polyclonal |
| Isotype: | IgG |
| Concentration: | 1 mg/ml |
| Observed Band: | 180kD |
| GeneID: | 2066 |
| Human Swiss-Prot No: | Q15303 |
| Cellular localization: | Cell membrane ; Single-pass type I membrane protein . In response to NRG1 treatment, the activated receptor is internalized.; [ERBB4 intracellular domain]: Nucleus . Mitochondrion . Following proteolytical processing E4ICD (E4ICD1 or E4ICD2 generated from the respective isoforms) is translocated to the nucleus. Significantly more E4ICD2 than E4ICD1 is found in the nucleus. E4ICD2 colocalizes with YAP1 in the nucleus. |
| Background: | This gene is a member of the Tyr protein kinase family and the epidermal growth factor receptor subfamily. It encodes a single-pass type I membrane protein with multiple cysteine rich domains, a transmembrane domain, a tyrosine kinase domain, a phosphotidylinositol-3 kinase binding site and a PDZ domain binding motif. The protein binds to and is activated by neuregulins and other factors and induces a variety of cellular responses including mitogenesis and differentiation. Multiple proteolytic events allow for the release of a cytoplasmic fragment and an extracellular fragment. Mutations in this gene have been associated with cancer. Alternatively spliced variants which encode different protein isoforms have been described; however, not all variants have been fully characterized. [provided by RefSeq, Jul 2008], |
-
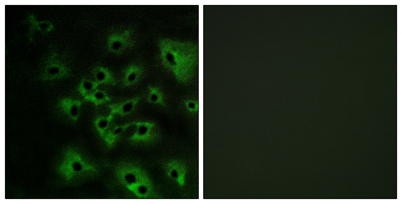
 Immunofluorescence analysis of HeLa cells treated with EGF 200nM 5', using HER4 (Phospho-Tyr1284) Antibody. The picture on the right is blocked with the phospho peptide.
Immunofluorescence analysis of HeLa cells treated with EGF 200nM 5', using HER4 (Phospho-Tyr1284) Antibody. The picture on the right is blocked with the phospho peptide. -
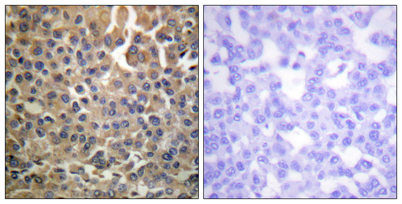
 Immunohistochemistry analysis of paraffin-embedded human breast carcinoma, using HER4 (Phospho-Tyr1284) Antibody. The picture on the right is blocked with the phospho peptide.
Immunohistochemistry analysis of paraffin-embedded human breast carcinoma, using HER4 (Phospho-Tyr1284) Antibody. The picture on the right is blocked with the phospho peptide. -
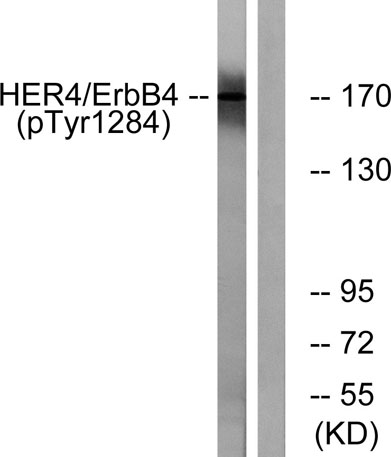
 Western blot analysis of lysates from HUVEC cells treated with EGF 200ng/ml 30', using HER4 (Phospho-Tyr1284) Antibody. The lane on the right is blocked with the phospho peptide.
Western blot analysis of lysates from HUVEC cells treated with EGF 200ng/ml 30', using HER4 (Phospho-Tyr1284) Antibody. The lane on the right is blocked with the phospho peptide.

 Manual
Manual